Mark O. Martin, PhD
My goal here in the Biology Department of the University of Puget Sound is to train students to approach problems in microbiology with critical thinking, enthusiasm, communication skills, more enthusiasm, and above all a microbiocentric point of view. The balance between teaching and research is, in my opinion, a false dichotomy: just as I teach in the classroom, I teach my research students at the bench. And it is true, in both situations, that I learn from my students as much as they learn from me! No matter the venue, mentoring, collaboration, creativity, and positivity are vital to me.
In the classroom, you can find me working with students in several courses here in Tacoma. I teach first year students an introductory lab based survey of cell and molecular biology (Biology 111, "The Unity of Life"). I teach a first year writing seminar focusing on symbiotic and parasitic relationships (SSI1 165, "Never Really Alone: Symbiosis and Parasitism Around and Within Us"); I also teach a lecture based version of this course to juniors and seniors (Biology 376 "ONE: Our Symbiotic Planet"). Finally, I teach a lab based overview of the depth and breadth of the microbial world to juniors and seniors (Biology 350, "Microbiology"---that yes, needs a snazzier title). In all of my courses, I focus on student-driven creative approaches and investigation in concert with more standard pedagogical approaches.

My undergraduate driven research lab focuses on what I term #MattersMicrobial. I have a background in microbial genetics, and in particular the genetics of unusual---"undomesticated"---microbes. In addition, I have a longstanding interest in microbial symbioses, which in turn has taken me far afield in more ecological approaches.
The areas of interest in my lab include bacterial predators, such as Bdellovibrio, Ensifer, and the enigmatic Saprospira. I also involved in an NSF-supported project (with Stacey Weiss in my department) to explore the possibility of cloacal microbes in the Striped Plateau Lizard acting as a protective strategy against fungal or bacterial attacks on newly laid eggs (I think of them as "microbial midwives").

Several of us are attempting to explore the microbiome of the charismatic and extremophilic tardigrade. I also work with Joel Elliott in my department to explore the microbial inhabitants of marine anthropogenic sulfide seeps here in Puget Sound. For many years, I have been interested in bacterial bioluminescence, which I use for artistic as well as scientific explorations. Finally, I am quite interested in outreach and science communication.
Thus, there is something for every #DocMartian to explore in the Microbial Menagerie!
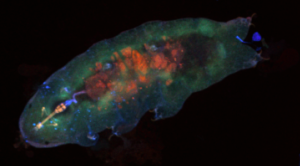
It is very important to me that each student in my laboratory has THEIR project. This slows progress, perhaps, but also yields the benefit of ownership and personal accountability. My approach works, in terms of the best outcomes: well prepared, thoughtful, and scientifically minded students.
I have been teaching and doing research at small liberal arts institutions since 1995 (I have been at the University of Puget Sound since 2005). I am extremely proud of all of my students, and find their points of view, insights, and positivity enriching. And my research students? I have sent 24 of them to PhD programs, and currently five are in faculty positions.

The students I have been fortunate enough to work with in my lab soon turn into interactive colleagues...and there is no better reward of the teacher-scholar model of education.
In my personal life, I am married to Dr. Jennifer Quinn, a mathematician at the University of Washington Tacoma (as well as a fabulous creative artist). We have two teenaged sons, Anson and Zachary. Now there are three eukaryotes of which I heartily approve!
References
- Bunker ME, Elliott G, Heyer-Gray H, Martin MO, Arnold AE, Weiss SL. “Vertically transmitted microbiome protects eggs from fungal infection and egg failure.” Anim Microbiome. 2021;3(1):43.
- Breitbart, M., Malki, K., Sawaya, N.A., Bonnain, C., and M.O. Martin (2018). “Elementary Student Outreach Activity Demonstrating the Use of ‘Phage Therapy Heroes’ to Combat Bacterial Infections.” J. Microbiol. Bio. Education. 19(1). pii: 19.1.30. doi: 10.1128/jmbe.v19i1.1407 pdf
- Williams, L.E., Baltrus, D.A., O’Donnell, S.D., Skelly, T.J., and M.O. Martin (2017). “Complete Genome Sequence of the Predatory Bacterium Ensifer adhaerens Casida A.” Genome Announc. 5: e01344-17. pdf
- Jetten, M.S.M, Martin, M.O., Romling, and V.J. Torres (2016). “The Endurance of Microbiology: An Interview With Mike Jetten, Mark Martin, Ute Romling, and Victor Torres.” Trends in Microbiology 24: 319 – 323.
- Martin, M. (2014). “Casting a Long Shadow in the Classroom: An Educator’s Perspective of the Contributions of Carl Woese.” RNA Biology: 11: 244 – 247.
- Shogan, BD, Smith, DP, Packman AI, Kelley, ST, Landon, EM, Bhangar, S., Vora, GJ, Jones, RM, Keegan, K., Stephens, B., Ramos, T., Kirkup, ST, Levin, H., Rosenthal, M., Foxman, B., Chang, EB, Siegel, J., Cobey, S., An, G., Alverdy, JC, Olsiewski, PJ, Martin, MO, Marrs, R., Hernandez, M, Christley, S., Morowitz, M., Weber, S., and J. Gilbert (2013). "The Hospital Microbiome Project: Meeting Report for the 2nd Hospital Microbiome Project, Chicago, USA, January 15th, 2013." Standards in Genomic Sciences 8: 571 – 579.
- Martin, M.O., Gilman*, F.R., and S.L. Weiss. (2010). “Sex-specific asymmetry within the cloacal microbiota of the striped plateau lizard, Sceloporus virgatus.” Symbiosis 51: 97 – 105.
- Megan E. Núñez, Mark O. Martin, Phyllis H. Chan*, Lin K. Duong*, Anil R. Sindhurakar*, and Eileen M. Spain. (2005). "Atomic Force Microscopy of Bacterial Communities." Methods in Enzymology, J. Leadbetter, editor 397: 256 – 268.
- Dunn, A.K., Martin, M.O. and E.V. Stabb. (2005). “Characterization of pES213, a small mobilizable plasmid from Vibrio fischeri.” Plasmid 54: 114 – 134.
- Nunez, M.E., Martin, M.O., Chan, P.H., and E.M. Spain. (2005). “Predation, death, and survival in a biofilm: Bdellovibrio investigated by atomic force microscopy.” Colloids Surf. B. Biointerfaces 42: 263 – 271.
- Nunez, M.E., Martin, M.O., Duong*, L.K., Ly*, E., and E.M. Spain. (2003). "Investigations into the life cycle of the bacterial predator Bdellovibrio bacteriovorus 109J at an interface by atomic force microscopy." Biophys. J. 84: 3379 - 3388.
- Martin, Mark O. (2002). "Predatory Prokaryotes: An Emerging Research Opportunity.” J. Mol. Microbiol. Biotechnol. 4: 467 - 477. pdf
- Martin, Mark O. (1995). “Biological conservation strategies: optimizing in situ and ex situ approaches.” Trends in Ecol. Evol. 10: 227 – 228.
- Martin, Mark O. (1994) “Synthesis of Commercially Valuable Bacterial Polymers: Impact of Molecular Genetics.” Res. Microbiol. 145: 93 - 97.
- Showalter, R., M. Martin, and M. Silverman. (1990). “Cloning and Nucleotide Sequence of lux R, a Regulatory Gene Controlling Bioluminescence in Vibrio harveyi.” J. Bacteriol. 172: 2946 - 2954.
- Martin, M., R. Showalter, and M. Silverman. (1990). “Identification of a Locus Controlling Expression of Luminescence Genes in Vibrio harveyi.” J. Bacteriol. 171: 2406 - 2414.
- Silverman, M., M. Martin, and J. Engebrecht. (1988). “Regulation of Luminescence in Marine Bacteria.” In: The Genetics of Bacterial Diversity, Academic Press, London, UK. D.A. Hopwood and K.F. Chater, editors.
- Martin, Mark O. and Sharon R. Long. (1984). “Generalized Transduction in Rhizobium meliloti.” J. Bacteriol. 159: 125 - 129.
- Russell Higuichi, Howard D. Stang, Jeffrey K. Browne, Mark O. Martin, Marie Huot, Janice Lipeles, and Winston Salser. (1981). “Human Ribosomal RNA Gene Spacer Sequences are Found Interspersed Elsewhere in the Genome.” Gene 15: 177 - 186.

